spectrex mock example#
End-to-end demonstration of the spectrex Phase 1 API using a synthetic (mock) scene on a 500 × 20 pixel NIRISS/GR150R grism stamp.
The pipeline:
Load instrument configuration and PCA eigenspectra basis
Build (or load) the sparse forward operator H
Generate a mock scene — sparse PCA coefficient vector ã
Forward-model: disperse ã through H to produce a grism image
Recover ã from the dispersed image by LSQR
Repeat with Poisson + read noise to demonstrate the realistic case
RMSE is computed on reconstructed flux values f(λ) = Φ @ a so it is
physically meaningful and matches the parity plot axes.
from pathlib import Path
import matplotlib.pyplot as plt
import numpy as np
import spectrex
from spectrex import (
EigenspectraBasis,
InstrumentConfig,
NoiseModel,
SciPySparseOperator,
SpectralSolver,
)
# ── Paths ────────────────────────────────────────────────────────────────────
TESTDATA = Path("../testdata")
OPERATOR_CACHE = Path("operator_cache.npz")
# ── Configuration ─────────────────────────────────────────────────────────────
COLD_START = False # set True to force operator rebuild from scratch
IMAGE_SHAPE = (500, 20)
SOURCE_DENSITY = 0.1 # fraction of pixels with injected sources
SEED = 42
N_COMPONENTS = 10 # must match eigenspectra CSV
rng = np.random.default_rng(SEED)
print(f"spectrex {spectrex.__version__}")
spectrex 0.2.1.dev18+gc6f5edaa6.d20260506
Step 1 — Instrument configuration & eigenspectra basis#
InstrumentConfig loads the aXe grism config file, wavelength range, and
sensitivity curves. EigenspectraBasis loads the 10 Kurucz PCA eigenspectra
and interpolates them onto the instrument wavelength grid.
config = InstrumentConfig.from_files(
conf_path=TESTDATA / "Config Files" / "GR150R.F150W.220725.conf",
wavelengthrange_path=TESTDATA / "jwst_niriss_wavelengthrange_0002.asdf",
sensitivity_dir=TESTDATA / "SenseConfig" / "wfss-grism-configuration",
filter_name="F150W",
n_wavelengths=150,
)
basis = EigenspectraBasis.from_csv(
TESTDATA / "eigenspectra_kurucz.csv",
config.wavelengths,
)
print(f"Wavelength range: {config.wavelengths[0]:.0f} – {config.wavelengths[-1]:.0f} Å")
print(f"Grism orders: {list(config.orders)}")
print(f"Basis components: {basis.n_components}")
Wavelength range: 12900 – 17100 Å
Grism orders: ['A', 'B', 'C', 'D', 'E']
Basis components: 10
Step 2 — Forward operator#
Builds the sparse matrix H (shape n_pix × n_pix·n_components) that maps
PCA coefficients to dispersed pixel values across all grism orders.
Note: The first run builds H from scratch, which takes ~60 s for a 500 × 20 stamp. The result is cached to
operator_cache.npz. SetCOLD_START = Truein the constants cell to force a rebuild.
import time
if OPERATOR_CACHE.exists() and not COLD_START:
op = SciPySparseOperator.load(OPERATOR_CACHE)
print(f"Operator loaded from {OPERATOR_CACHE} "
f"(shape {op._H.shape[0]} × {op._H.shape[1]})")
else:
t0 = time.perf_counter()
op = SciPySparseOperator.build(config, basis, IMAGE_SHAPE)
op.save(OPERATOR_CACHE)
elapsed = time.perf_counter() - t0
print(f"Operator built in {elapsed:.1f} s — cached to {OPERATOR_CACHE}")
print(f"Shape: {op._H.shape[0]} × {op._H.shape[1]}")
Operator loaded from operator_cache.npz (shape 10000 × 100000)
Step 3 — Mock scene#
We inject sources at a random 10 % of pixels. For each active pixel we draw random PCA coefficients, accept only if the reconstructed spectrum is non-negative (physical), and store them in the flat coefficient vector ã.
The direct image is the broadband integrated flux I(x,y) = ∫ a(x,y,λ) dλ, visualised as a 2-D stamp.
n_pix = IMAGE_SHAPE[0] * IMAGE_SHAPE[1]
n = basis.n_components
a_tilde = np.zeros(n_pix * n)
num_active = int(SOURCE_DENSITY * n_pix)
active_k = rng.choice(n_pix, size=num_active, replace=False)
MAX_TRIES = 50
n_placed = 0
for k in active_k:
for _ in range(MAX_TRIES):
flux = rng.uniform(-1, 1, size=n)
if np.all(basis.reconstruct(flux) >= 0):
a_tilde[k * n : (k + 1) * n] = flux
n_placed += 1
break
print(f"Sources placed: {n_placed} / {num_active} requested "
f"({100 * n_placed / n_pix:.1f} % of pixels)")
direct = basis.broadband_image(a_tilde, IMAGE_SHAPE)
print(f"Direct image shape: {direct.shape}, max flux: {direct.max():.4f}")
Sources placed: 896 / 1000 requested (9.0 % of pixels)
Direct image shape: (500, 20), max flux: 698067752.2096
Noiseless recovery#
Forward-model ã → dispersed grism image, then recover ã from the dispersed image using LSQR. The support mask is built from the direct image (non-zero pixels) so LSQR only solves for active columns of H.
dispersed = op.apply(a_tilde).reshape(IMAGE_SHAPE)
# Support mask: True at pixels known to have a source
support_mask = a_tilde != 0
print(f"Dispersed image range: [{dispersed.min():.4f}, {dispersed.max():.4f}]")
print(f"Active coefficients: {support_mask.sum()} / {len(support_mask)}")
Dispersed image range: [0.0000, 17101.3035]
Active coefficients: 8960 / 100000
solver = SpectralSolver(op, max_iter=500, tolerance=1e-8)
recovered = solver.solve(dispersed, support_mask=support_mask)
print(f"Recovered vector shape: {recovered.shape}")
Recovered vector shape: (100000,)
active_indices = [k for k in range(n_pix) if np.any(a_tilde[k * n : (k + 1) * n] != 0)]
true_flux = np.concatenate(
[basis.reconstruct(a_tilde[k * n : (k + 1) * n]) for k in active_indices]
)
rec_flux = np.concatenate(
[basis.reconstruct(recovered[k * n : (k + 1) * n]) for k in active_indices]
)
rmse_noiseless = np.sqrt(np.mean((true_flux - rec_flux) ** 2))
print(f"Noiseless RMSE (flux): {rmse_noiseless:.6f}")
Noiseless RMSE (flux): 21287.860870
def _clip(arr, nsigma_lo=2, nsigma_hi=2):
m, s = np.nanmean(arr), np.nanstd(arr)
return m - nsigma_lo * s, m + nsigma_hi * s
recovered_img = basis.broadband_image(recovered, IMAGE_SHAPE)
residual_img = np.abs(direct - recovered_img)
vmin_dr, vmax_dr = _clip(direct) # shared scale for Direct & Recovered
vmin_d2, vmax_d2 = _clip(dispersed)
vmax_res = np.nanmean(residual_img) + np.nanstd(residual_img)
fig, axes = plt.subplots(1, 4, figsize=(14, 4))
kw = dict(origin="lower", aspect="auto", interpolation="nearest", cmap="inferno")
im0 = axes[0].imshow(direct, vmin=vmin_dr, vmax=vmax_dr, **kw)
im1 = axes[1].imshow(dispersed, vmin=vmin_d2, vmax=vmax_d2, **kw)
im2 = axes[2].imshow(recovered_img, vmin=vmin_dr, vmax=vmax_dr, **kw)
im3 = axes[3].imshow(residual_img, vmin=0, vmax=vmax_res, **kw)
titles = ["Direct image", "Dispersed (grism)", "Recovered", "|Residual|"]
for ax, im, title in zip(axes, [im0, im1, im2, im3], titles):
ax.set_title(title)
ax.set_xlabel("column")
ax.set_ylabel("row")
fig.colorbar(im, ax=ax, fraction=0.046, pad=0.04)
fig.suptitle("Noiseless recovery — 500 × 20 stamp, GR150R/F150W", y=1.01)
fig.tight_layout()
plt.show()
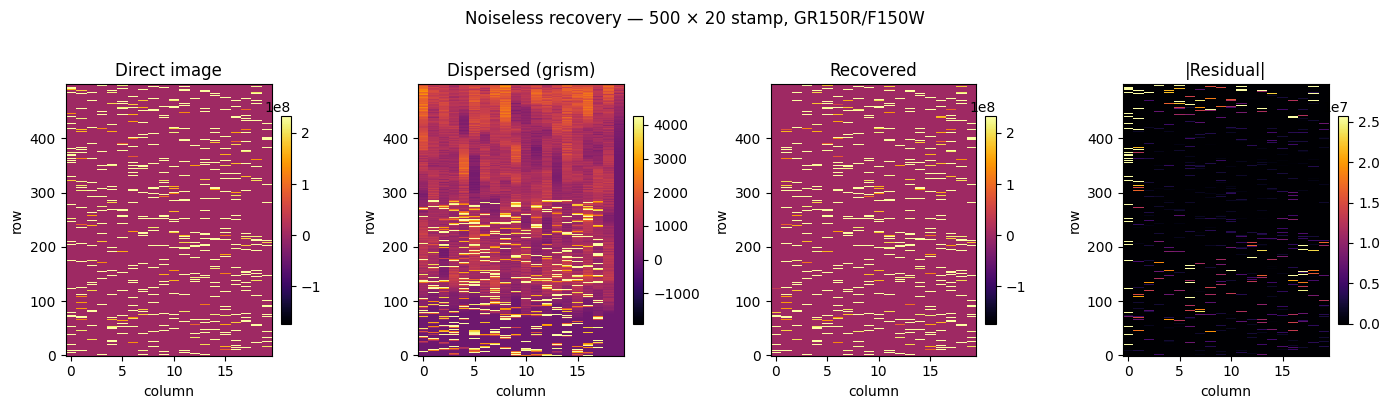
The dispersion process includes sources that do not reside entirely within the dispersed image due to positional shifts. This is an edge effect that we should include in a future version by virtually extending the pixel coordinates of the model image to negative and beyond the original size values in the dispersion direction.
fig, axes = plt.subplots(2, 1, figsize=(5, 8), sharex=True, tight_layout=True, height_ratios=(1, 0.6), gridspec_kw={'hspace': 0})
ax = axes[0]
outliers = rec_flux <= 1.
ax.scatter(true_flux[~outliers], rec_flux[~outliers], s=2, alpha=0.05, linewidths=0, color="C0", rasterized=True)
ax.scatter(true_flux[outliers], rec_flux[outliers], s=2, alpha=0.05, linewidths=0, color="C1", rasterized=True)
minv = max(min(true_flux.min(), rec_flux.min()), -10_000)
maxv = max(true_flux.max(), rec_flux.max())
ax.plot([minv, maxv], [minv, maxv], "r--", lw=1, label="1:1")
ax.set_aspect("equal", adjustable="box")
ax.set_xlim(minv, maxv)
ax.set_ylim(minv, maxv)
ax.set_ylabel("Recovered flux f(λ)")
ax.set_title("Parity plot — noiseless")
ax = axes[1]
frac_residuals = (true_flux - rec_flux) / true_flux
ax.scatter(true_flux[~outliers], frac_residuals[~outliers], s=2, alpha=0.05, linewidths=0, color="C0", rasterized=True)
ax.scatter(true_flux[outliers], frac_residuals[outliers], s=2, alpha=0.05, linewidths=0, color="C1", rasterized=True)
ax.set_ylim(-2, 2)
ax.set_xlabel("True flux f(λ)")
rmse_noiseless_good = np.sqrt(np.mean(frac_residuals[~outliers]) ** 2)
ax.text(0.05, 0.92, f"frac. mean error = {rmse_noiseless_good:.4f}", color='C0',
transform=ax.transAxes, fontsize=10,
bbox=dict(boxstyle="round,pad=0.3", fc="white", ec="gray", alpha=0.8))
fig.tight_layout()
plt.show()
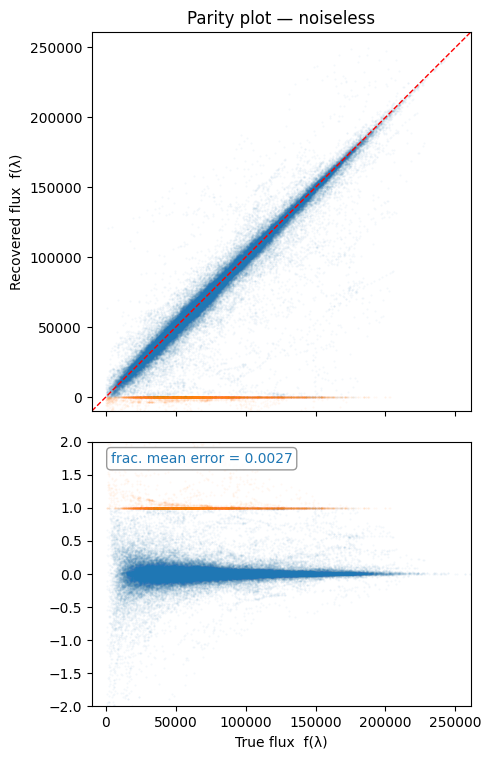
For this plot we highlight the failed recovery due to mapping coverage mentioned above. The fractional mean error is calculated without these outliers.
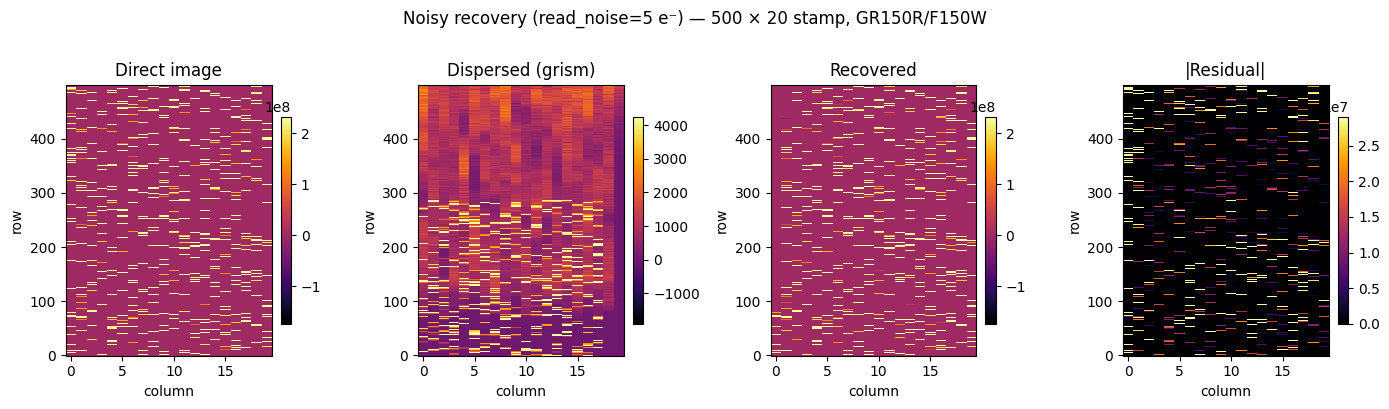
Noisy recovery#
We add Gaussian noise approximating Poisson + detector read noise
(read_noise = 5 e⁻) using NoiseModel.sample(). The same support mask
and LSQR solver are used; the only change is the input image.
This is the realistic operating regime for real JWST data.
noise_model = NoiseModel(read_noise=5.0)
noisy_dispersed = noise_model.sample(dispersed, rng)
print(f"Noisy dispersed range: [{noisy_dispersed.min():.4f}, {noisy_dispersed.max():.4f}]")
print(f"Added noise std (mean over pixels): "
f"{np.std(noisy_dispersed - dispersed):.4f}")
Noisy dispersed range: [-24.6412, 16906.7927]
Added noise std (mean over pixels): 34.1373
noisy_recovered = solver.solve(noisy_dispersed, support_mask=support_mask)
print(f"Noisy recovered vector shape: {noisy_recovered.shape}")
Noisy recovered vector shape: (100000,)
noisy_rec_flux = np.concatenate(
[basis.reconstruct(noisy_recovered[k * n : (k + 1) * n]) for k in active_indices]
)
rmse_noisy = np.sqrt(np.mean((true_flux - noisy_rec_flux) ** 2))
print(f"Noiseless RMSE: {rmse_noiseless:.6f}")
print(f"Noisy RMSE: {rmse_noisy:.6f}")
print(f"RMSE ratio: {rmse_noisy / rmse_noiseless:.2f}×")
Noiseless RMSE: 21287.860870
Noisy RMSE: 25840.669446
RMSE ratio: 1.21×
noisy_recovered_img = basis.broadband_image(noisy_recovered, IMAGE_SHAPE)
noisy_residual_img = np.abs(direct - noisy_recovered_img)
vmax_nres = np.nanmean(noisy_residual_img) + np.nanstd(noisy_residual_img)
fig, axes = plt.subplots(1, 4, figsize=(14, 4))
im0 = axes[0].imshow(direct, vmin=vmin_dr, vmax=vmax_dr, **kw)
im1 = axes[1].imshow(noisy_dispersed, vmin=vmin_d2, vmax=vmax_d2, **kw)
im2 = axes[2].imshow(noisy_recovered_img,vmin=vmin_dr, vmax=vmax_dr, **kw)
im3 = axes[3].imshow(noisy_residual_img, vmin=0, vmax=vmax_nres, **kw)
for ax, im, title in zip(axes, [im0, im1, im2, im3], titles):
ax.set_title(title)
ax.set_xlabel("column")
ax.set_ylabel("row")
fig.colorbar(im, ax=ax, fraction=0.046, pad=0.04)
fig.suptitle("Noisy recovery (read_noise=5 e⁻) — 500 × 20 stamp, GR150R/F150W", y=1.01)
fig.tight_layout()
plt.show()
fig, axes = plt.subplots(2, 1, figsize=(5, 8), sharex=True, tight_layout=True, height_ratios=(1, 0.6), gridspec_kw={'hspace': 0})
ax = axes[0]
outliers = noisy_rec_flux <= 1.
minv = max(min(true_flux.min(), noisy_rec_flux.min()), -10_000)
maxv = max(true_flux.max(), noisy_rec_flux.max())
ax.scatter(true_flux[~outliers], noisy_rec_flux[~outliers], s=2, alpha=0.05, linewidths=0, color="C0", rasterized=True)
ax.scatter(true_flux[outliers], noisy_rec_flux[outliers], s=2, alpha=0.05, linewidths=0, color="C1", rasterized=True)
minv = max(min(true_flux.min(), noisy_rec_flux.min()), -10_000)
maxv = max(true_flux.max(), noisy_rec_flux.max())
ax.plot([minv, maxv], [minv, maxv], "r--", lw=1, label="1:1")
ax.set_aspect("equal", adjustable="box")
ax.set_xlim(minv, maxv)
ax.set_ylim(minv, maxv)
ax.set_ylabel("Recovered flux f(λ)")
ax.set_title("Parity plot — noiseless")
ax = axes[1]
frac_residuals = (true_flux - noisy_rec_flux) / true_flux
ax.scatter(true_flux[~outliers], frac_residuals[~outliers], s=2, alpha=0.05, linewidths=0, color="C0", rasterized=True)
ax.scatter(true_flux[outliers], frac_residuals[outliers], s=2, alpha=0.05, linewidths=0, color="C1", rasterized=True)
ax.set_ylim(-2, 2)
ax.set_xlabel("True flux f(λ)")
rmse_noiseless_good = np.sqrt(np.mean(frac_residuals[~outliers]) ** 2)
ax.text(0.05, 0.92, f"frac. mean error = {rmse_noiseless_good:.4f}", color='C0',
transform=ax.transAxes, fontsize=10,
bbox=dict(boxstyle="round,pad=0.3", fc="white", ec="gray", alpha=0.8))
fig.tight_layout()
plt.show()
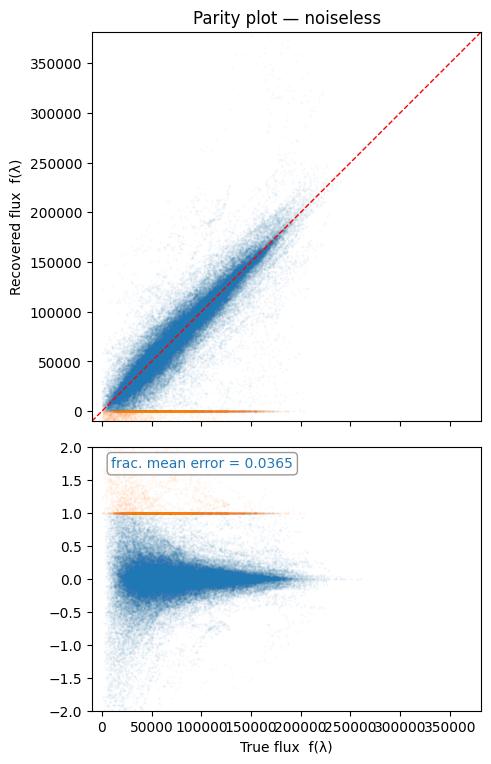
Next steps#
The parity plots above give a single RMSE figure for one source density. To quantify how recovery quality degrades as the field becomes more crowded, see the planned sweep notebook:
notebooks/analysis_rmse_vs_density.ipynb
That notebook will define a run_pipeline(a_tilde, op, basis, noise_model=None)
helper and sweep SOURCE_DENSITY over ~5–8 values, plotting RMSE vs density
for both the noiseless and noisy cases. This sweep is important for
establishing the credibility of the method in crowded NIRISS fields.